【源码阅读】SIR-GN: A Fast Structural Iterative Representation Learnin
@TOC

简介
Hello!
非常感谢您阅读海轰的文章,倘若文中有错误的地方,欢迎您指出~
ଘ(੭ˊᵕˋ)੭
昵称:海轰
标签:程序猿|C++选手|学生
简介:因C语言结识编程,随后转入计算机专业,获得过国家奖学金,有幸在竞赛中拿过一些国奖、省奖…已保研
学习经验:扎实基础 + 多做笔记 + 多敲代码 + 多思考 + 学好英语!
唯有努力💪
论文简介
原文链接:https://dl.acm.org/doi/abs/10.1145/3450315
期刊:ACM Transactions on Knowledge Discovery from Data (CCF B类)
源码:https://github.com/mjoaristi/SIR-GN
年度:2021/05/19
准备工作
环境
- python3.7
- numpy
- sklearn
创建一个python 3.7的环境 再安装numpy、sklearn两个包就好了
再导入Pycharm,选择刚刚创建的环境,设置好工作目录就好了
运行
找到run_sirgn.sh文件

运行即可(前提得先设置工作路径)
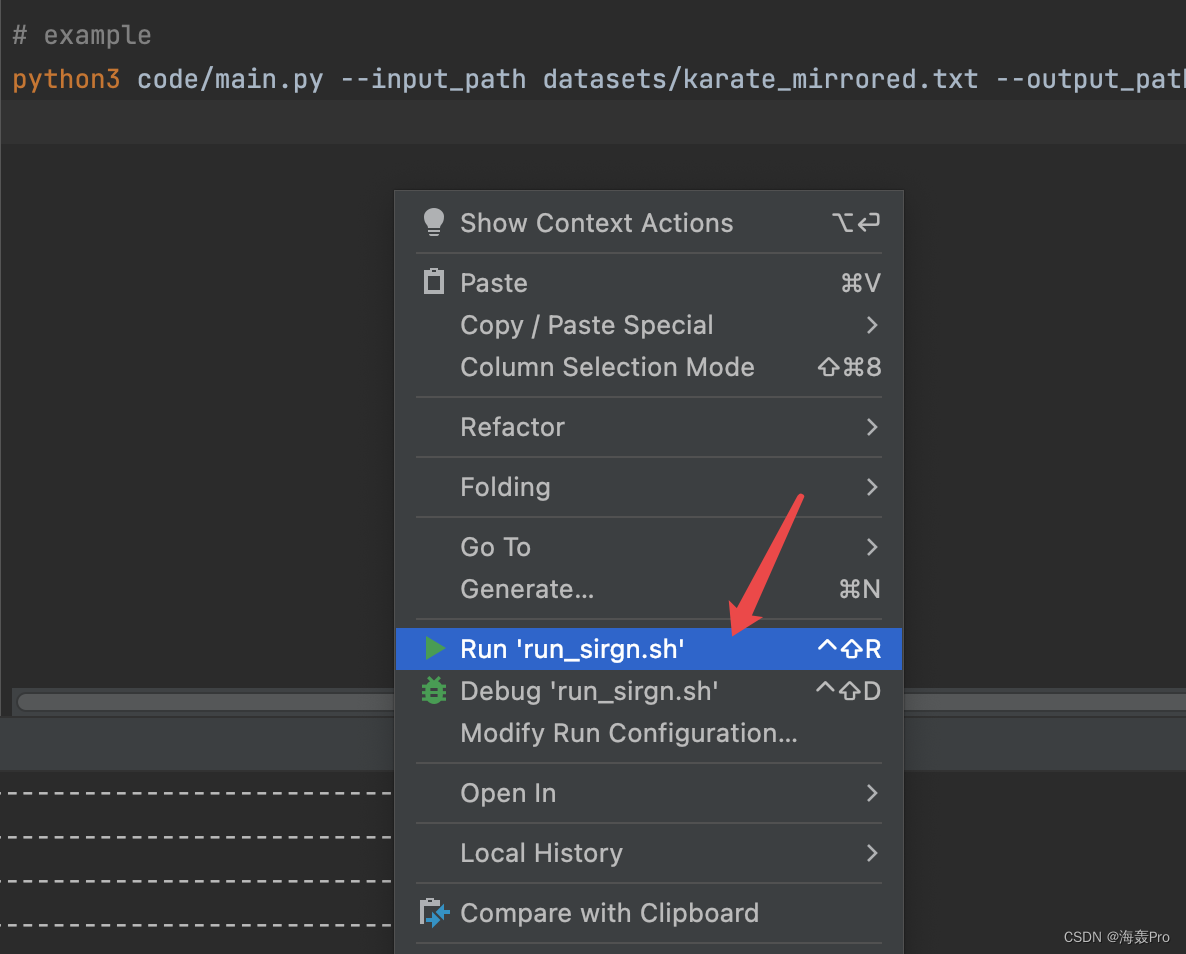
运行结果
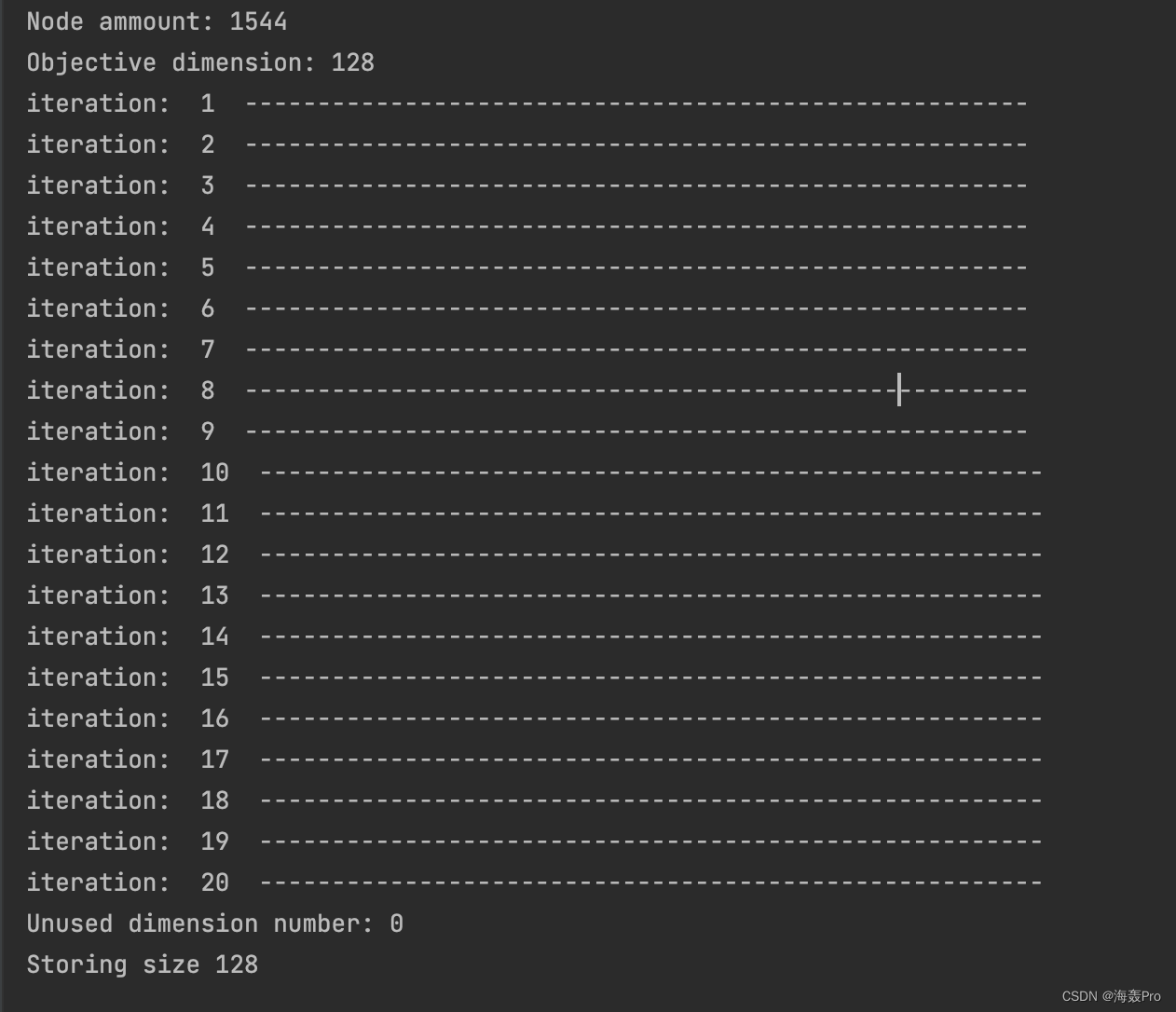
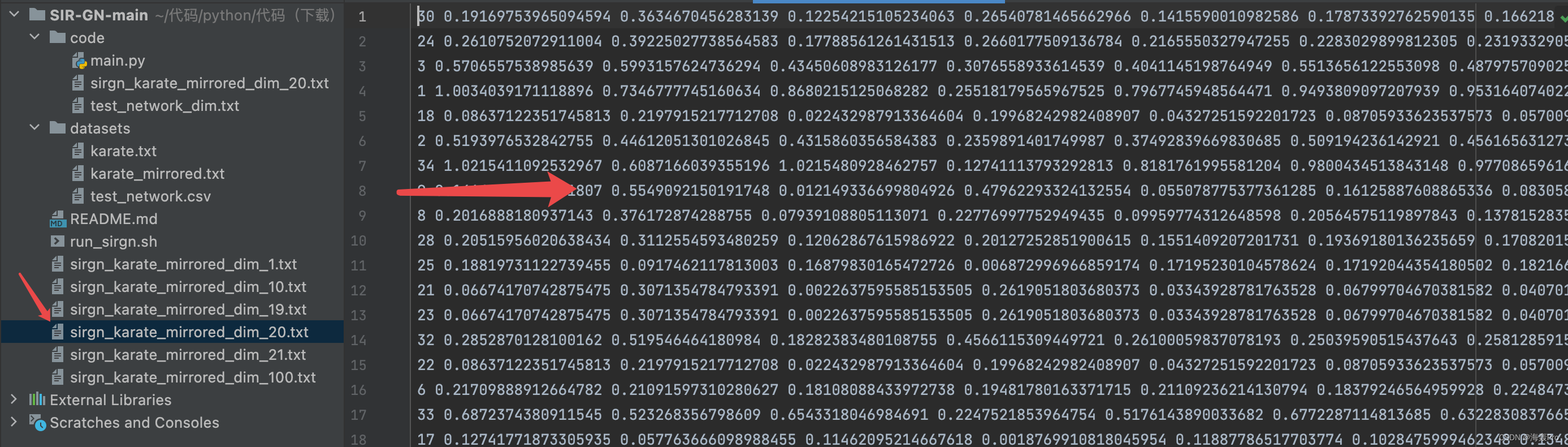
源码阅读
参数设置
- –input_path:图存放的路径
- –embedding_size:最终嵌入的大小
- –max_it:迭代次数
- –output_path:最终生成嵌入文件存放的位置
args = parse_arguments()
# parse_arguments()
def parse_arguments():
parser = argparse.ArgumentParser(description="Run SIR-GN.")
parser.add_argument('--input_path',default='../datasets/test_network.csv', type = str, action="store", dest="input_path", help='Input graph path.')
parser.add_argument('--embedding_size', type = int, default= 128,action="store", dest="embedding_size", help='Embedding size.')
parser.add_argument('--max_it', type = int, default= 20, action="store", dest="max_it", help='Maximum amount of iterations.')
parser.add_argument('--output_path', type = str, default='./test_network_dim.txt', action="store", dest="output_path", help='Output embedding file path.')
return parser.parse_args()
初始化类SirGN
p = SirGN(args.input_path,embsize=args.embedding_size,maxIter=args.max_it)
类初始化
- 读取图文件(存储节点与节点的对应关系)中所有存在的节点
self.nodes - 同时记录每个节点的邻域集合
self.outedge - 最后生成节点的初始嵌入
self.sol
def __init__(self, st,embsize=50,maxIter=1000,sep=','):
# st: 图文件路经
self.embsize=embsize
self.maxIter=maxIter
self.nodes=dict() # 节点
self.outedge=dict() # 节点对应的邻域节点
self.cont=0 # 节点个数
with open(st, "r", encoding = "utf-8") as c:
for l in c:
a=l.split(sep)
b=(int)(a[0].replace('\n',''))
a=(int)(a[1].replace('\n',''))
if not a in self.nodes:
self.nodes[a]=self.cont
self.cont=self.cont+1
if not self.nodes[a] in self.outedge:
self.outedge[self.nodes[a]]=set()
if not b in self.nodes:
self.nodes[b]=self.cont
self.cont=self.cont+1
if not self.nodes[b] in self.outedge:
self.outedge[self.nodes[b]]=set()
with open(st, "r", encoding = "utf-8") as c:
for l in c:
a=l.split(sep)
# b=a[1].replace('\n','')
# a=a[0].replace('\n','')
b=(int)(a[0].replace('\n',''))
a=(int)(a[1].replace('\n',''))
self.outedge[self.nodes[a]].add(self.nodes[b])
self.outedge[self.nodes[b]].add(self.nodes[a])
# sol:节点对应的嵌入(开始的时候初始化,第一列为节点的度,其余为0)
self.sol=self.initialize()
这里使用
cont大概意思就是重新对节点进行编号
例如,图文档中节点是35号,但是其是第一个读入的,编号为1
估计作者这样写的动机是防止初始节点编号的稀疏性,避免浪费后面需要开创的空间
初始化最开始的节点嵌入(shape = n * embsize)
- 第一列为节点对应的度
- 其余列为0
def initialize(self):
# dimr 每个节点对应的度
dimr=[len(self.outedge[x]) for x in range(self.cont)]
print('Node ammount: {}'.format(len(dimr)))
dim_list = list(set(dimr)) # 去重 再转list
dim_list.sort() # 排序
dist_dim = len(dim_list)
dim_to_pos = dict(zip(dim_list,range(dist_dim)))
print('Objective dimension: {}'.format(self.embsize))
if self.embsize > len(dimr):
print('Objective dimension bigger that node ammount')
print('New objective dimension: {}'.format(len(dimr)))
self.embsize = len(dimr)
ret = np.zeros((self.cont,self.embsize))
# 第一次初始化 第0列为节点的度
for i,d in enumerate(dimr):
ret[i,0]=d
# cluster_labels:标签
self.cluster_labels = [dim_to_pos[x] for x in dimr]
# ret:初始嵌入
return ret
1.
dimr是度列表,每一个节点对应一个度
2.dim_list,用于记录度的不同数量(去重后),可以用来查看节点度的分布情况,当然这里是用于初始节点对应的标签(按照最开始的度划分就可以了)
3.为什么会有判断if self.embsize > len(dimr):?
因为embsize不仅代表嵌入维度,也还有簇种类的原因,n个节点,最多可以划分为n个簇,当节点总数小于簇总数时,不能保证每个簇都有节点,所有这里进行了一次判断,当节点数小于簇数量时,就令嵌入数等于节点数
开始进行迭代学习
p.learn()
这里的算法步骤与论文中完全一致(以k-means为例)
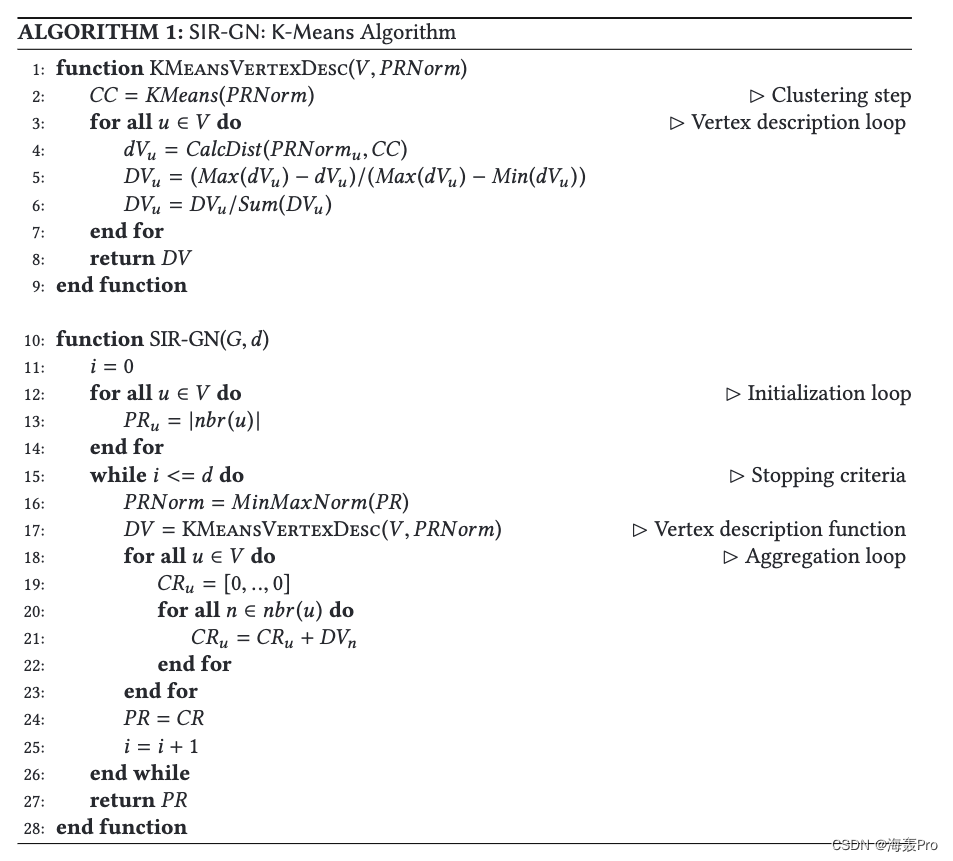

def learn(self):
it1=0
scaler = MinMaxScaler()
# sol:最开始初始化的那个矩阵(第0列为度 其余为0)
self.sol = scaler.fit_transform(self.sol)
while it1<self.maxIter:
print('iteration: ',it1+1,' ------------------------------------------------------')
scaler = MinMaxScaler()
self.sol = scaler.fit_transform(self.sol)
# print(self.sol.shape[1])
cluster_alg = KMeans(n_clusters=self.sol.shape[1],random_state=100)
# 更新labels
self.cluster_labels = cluster_alg.fit_predict(self.sol)
t = cluster_alg.fit_predict(self.sol)
used_set = set(self.cluster_labels)
dist_mat = np.zeros_like(self.sol) # size = n * embsize
# 依次遍历节点
for p1 in range(self.sol.shape[0]):
max_dist= 0 # 记录最大距离
min_dist= 1000000000 # 距离最小距离
dist_dict= {}
# 依次遍历每个质心
for p2 in range(cluster_alg.cluster_centers_.shape[0]):
if p2 in used_set:
# np.linalg.norm: 求范数 默认 2范数
# 计算每一个节点 与 每一个质心之间的距离
current_dist = np.linalg.norm(self.sol[p1,:]-cluster_alg.cluster_centers_[p2,:])
dist_dict[p2] = current_dist
if current_dist > max_dist:
max_dist = current_dist
if current_dist < min_dist:
min_dist = current_dist
# 按照离质心之间的距离 计算属于此簇的...(利用最大/最小距离)
# 没有出现过的簇 概率为0
for p2 in range(cluster_alg.cluster_centers_.shape[0]):
if p2 in used_set:
dist_mat[p1,p2] = (max_dist-dist_dict[p2])/(max_dist-min_dist)
else:
dist_mat[p1,p2] = 0
# 归一化 范围 0-1
for i_ in range(self.sol.shape[0]):
dist_mat[i_,:]/=np.sum(dist_mat[i_,:])
# generate emb
ret = np.zeros_like(self.sol)
for i in range(self.sol.shape[0]):
for neigh_id in self.outedge[i]:
ret[i,:]+=dist_mat[neigh_id,:]
self.sol= ret
it1+=1
最后写入嵌入,算法完成
p.write(args.output_path)
# 写入
def write(self,f):
if np.max(self.cluster_labels) < self.embsize:
print('Unused dimension number: {}'.format(self.embsize-(np.max(self.cluster_labels)+1)))
out_sol = self.sol
print('Storing size {}'.format(out_sol.shape[1]))
file1 = open(f,'w')
for h in self.nodes:
l=self.nodes[h]
v=''
ll=out_sol[l,:]
for g in range(ll.shape[0]):
v+=' '+str(ll[g])
file1.write(str(h)+v+'\n')
file1.close()
完整源码(注释版)
注:对比原版本,修改了一些地方,主要是测试了一下其他数据集(数据读入那一块)
import argparse
import numpy as np
from sklearn.preprocessing import MinMaxScaler
from sklearn.cluster import KMeans
def parse_arguments():
parser = argparse.ArgumentParser(description="Run SIR-GN.")
parser.add_argument('--input_path',default='../datasets/test_network.csv', type = str, action="store", dest="input_path", help='Input graph path.')
parser.add_argument('--embedding_size', type = int, default= 128,action="store", dest="embedding_size", help='Embedding size.')
parser.add_argument('--max_it', type = int, default= 20, action="store", dest="max_it", help='Maximum amount of iterations.')
parser.add_argument('--output_path', type = str, default='./test_network_dim.txt', action="store", dest="output_path", help='Output embedding file path.')
return parser.parse_args()
class SirGN:
def __init__(self, st,embsize=50,maxIter=1000,sep=','):
# st: 图文件路经
self.embsize=embsize
self.maxIter=maxIter
self.nodes=dict() # 节点
self.outedge=dict() # 节点对应的邻域节点
self.cont=0 # 节点个数
with open(st, "r", encoding = "utf-8") as c:
for l in c:
a=l.split(sep)
b=(int)(a[0].replace('\n',''))
a=(int)(a[1].replace('\n',''))
if not a in self.nodes:
self.nodes[a]=self.cont
self.cont=self.cont+1
if not self.nodes[a] in self.outedge:
self.outedge[self.nodes[a]]=set()
if not b in self.nodes:
self.nodes[b]=self.cont
self.cont=self.cont+1
if not self.nodes[b] in self.outedge:
self.outedge[self.nodes[b]]=set()
with open(st, "r", encoding = "utf-8") as c:
for l in c:
a=l.split(sep)
# b=a[1].replace('\n','')
# a=a[0].replace('\n','')
b=(int)(a[0].replace('\n',''))
a=(int)(a[1].replace('\n',''))
self.outedge[self.nodes[a]].add(self.nodes[b])
self.outedge[self.nodes[b]].add(self.nodes[a])
self.sol=self.initialize()
def initialize(self):
# dimr 每个节点对应的度
dimr=[len(self.outedge[x]) for x in range(self.cont)]
print('Node ammount: {}'.format(len(dimr)))
dim_list = list(set(dimr)) # 去重 再转list
dim_list.sort() # 排序
dist_dim = len(dim_list)
dim_to_pos = dict(zip(dim_list,range(dist_dim)))
print('Objective dimension: {}'.format(self.embsize))
if self.embsize > len(dimr):
print('Objective dimension bigger that node ammount')
print('New objective dimension: {}'.format(len(dimr)))
self.embsize = len(dimr)
ret = np.zeros((self.cont,self.embsize))
# 第一次初始化 第0列为节点的度
for i,d in enumerate(dimr):
ret[i,0]=d
# cluster_labels:标签
self.cluster_labels = [dim_to_pos[x] for x in dimr]
return ret
def learn(self):
it1=0
scaler = MinMaxScaler()
# sol:最开始初始化的那个矩阵(第0列为度 其余为0)
self.sol = scaler.fit_transform(self.sol)
while it1<self.maxIter:
print('iteration: ',it1+1,' ------------------------------------------------------')
scaler = MinMaxScaler()
self.sol = scaler.fit_transform(self.sol)
# print(self.sol.shape[1])
cluster_alg = KMeans(n_clusters=self.sol.shape[1],random_state=100)
# 更新labels
self.cluster_labels = cluster_alg.fit_predict(self.sol)
t = cluster_alg.fit_predict(self.sol)
used_set = set(self.cluster_labels)
dist_mat = np.zeros_like(self.sol) # size = n * embsize
# 依次遍历节点
for p1 in range(self.sol.shape[0]):
max_dist= 0 # 记录最大距离
min_dist= 1000000000 # 距离最小距离
dist_dict= {}
# 依次遍历每个质心
for p2 in range(cluster_alg.cluster_centers_.shape[0]):
if p2 in used_set:
# np.linalg.norm: 求范数 默认 2范数
# 计算每一个节点 与 每一个质心之间的距离
current_dist = np.linalg.norm(self.sol[p1,:]-cluster_alg.cluster_centers_[p2,:])
dist_dict[p2] = current_dist
if current_dist > max_dist:
max_dist = current_dist
if current_dist < min_dist:
min_dist = current_dist
# 按照离质心之间的距离 计算属于此簇的...(利用最大/最小距离)
# 没有出现过的簇 概率为0
for p2 in range(cluster_alg.cluster_centers_.shape[0]):
if p2 in used_set:
dist_mat[p1,p2] = (max_dist-dist_dict[p2])/(max_dist-min_dist)
else:
dist_mat[p1,p2] = 0
# 归一化 范围 0-1
for i_ in range(self.sol.shape[0]):
dist_mat[i_,:]/=np.sum(dist_mat[i_,:])
# generate emb
ret = np.zeros_like(self.sol)
for i in range(self.sol.shape[0]):
for neigh_id in self.outedge[i]:
ret[i,:]+=dist_mat[neigh_id,:]
self.sol= ret
it1+=1
def write(self,f):
if np.max(self.cluster_labels) < self.embsize:
print('Unused dimension number: {}'.format(self.embsize-(np.max(self.cluster_labels)+1)))
out_sol = self.sol
print('Storing size {}'.format(out_sol.shape[1]))
file1 = open(f,'w')
for h in self.nodes:
l=self.nodes[h]
v=''
ll=out_sol[l,:]
for g in range(ll.shape[0]):
v+=' '+str(ll[g])
file1.write(str(h)+v+'\n')
file1.close()
args = parse_arguments()
p = SirGN(args.input_path,embsize=args.embedding_size,maxIter=args.max_it)
p.learn()
p.write(args.output_path)
结语
文章仅作为个人学习笔记记录,记录从0到1的一个过程
希望对您有一点点帮助,如有错误欢迎小伙伴指正

- 点赞
- 收藏
- 关注作者

评论(0)